Insider Brief
- Cleveland Clinic and IBM researchers are laying the groundwork for applying quantum computing methods to protein structure prediction.
- The team’s combination of classical and quantum computing methods is an essential step for advancing our understanding of protein structures
- The team published their findings in the Journal of Chemical Theory and Computation.
PRESS RELEASE — Researchers from Cleveland Clinic and IBM recently published findings in the Journal of Chemical Theory and Computation that could lay the groundwork for applying quantum computing methods to protein structure prediction. This publication is the first peer-reviewed quantum computing paper from the Cleveland Clinic-IBM Discovery Accelerator partnership.
For decades, researchers have leveraged computational approaches to predict protein structures. A protein folds itself into a structure that determines how it functions and binds to other molecules in the body. These structures determine many aspects of human health and disease.
By accurately predicting the structure of a protein, researchers can better understand how diseases spread and thus how to develop effective therapies. Cleveland Clinic postdoctoral fellow Bryan Raubenolt, Ph.D., and IBM researcher Hakan Doga, Ph.D., spearheaded a team to discover how quantum computing can improve current methods.

In recent years, machine learning techniques have made significant progress in protein structure prediction. These methods are reliant on training data (a database of experimentally determined protein structures) to make predictions. This means that they are constrained by how many proteins they have been taught to recognize. This can lead to lower levels of accuracy when the programs/algorithms encounter a protein that is mutated or very different from those on which they were trained, which is common with genetic disorders.
The alternative method is to simulate the physics of protein folding. Simulations allow researchers to look at a given protein’s various possible shapes and find the most stable one. The most stable shape is critical for drug design.
The challenge is that these simulations are nearly impossible on a classical computer, beyond a certain protein size. In a way, increasing the size of the target protein is comparable to increasing the dimensions of a Rubik’s cube. For a small protein with 100 amino acids, a classical computer would need the time equal to the age of the universe to exhaustively search all the possible outcomes, says Dr. Raubenolt.
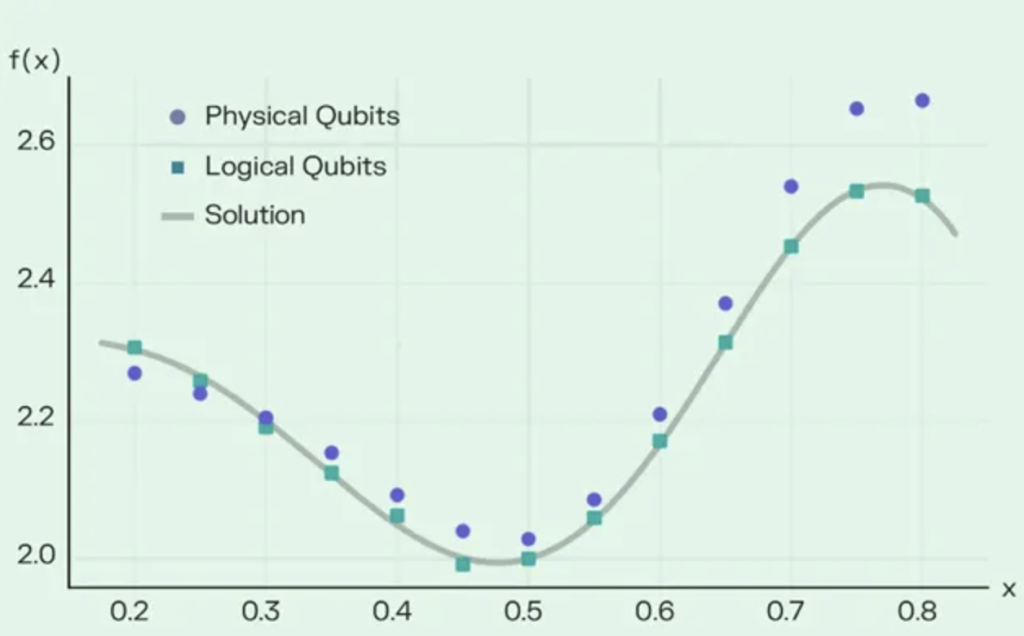
To help overcome these limitations, the research team applied a mix of quantum and classical computing methods. This framework could allow quantum algorithms to address the areas that are challenging for state-of-the-art classical computing, including protein size, intrinsic disorder, mutations and the physics involved in proteins folding. The framework was validated by accurately predicting the folding of a small fragment of a Zika virus protein on a quantum computer, compared to state-of-the-art classical methods.
The quantum-classical hybrid framework’s initial results outperformed both a classical physics-based method and AlphaFold2. Although the latter is designed to work best with larger proteins, it nonetheless demonstrates this framework’s ability to create accurate models without directly relying on substantial training data.
The researchers used a quantum algorithm to first model the lowest energy conformation for the fragment’s backbone, which is typically the most computationally demanding step of the calculation. Classical approaches were then used to convert the results obtained from the quantum computer, reconstruct the protein with its sidechains, and perform final refinement of the structure with classical molecular mechanics force fields. The project shows one of the ways that problems can be deconstructed into parts, with quantum computing methods addressing some parts and classical computing others, for increased accuracy.
Multidisciplinary collaboration was essential to achieve this framework.
“One of the most unique things about this project is the number of disciplines involved,” says Dr. Raubenolt. “Our team’s expertise ranges from computational biology and chemistry, structural biology, software and automation engineering, to experimental atomic and nuclear physics, mathematics, and of course quantum computing and algorithm design. It took the knowledge from each of these areas to create a computational framework that can mimic one of the most important processes for human life.”
The team’s combination of classical and quantum computing methods is an essential step for advancing our understanding of protein structures, and how they impact our ability to treat and prevent disease. The team plans to continue developing and optimizing quantum algorithms that can predict the structure of larger and more sophisticated proteins.
“This work is an important step forward in exploring where quantum computing capabilities could show strengths in protein structure prediction,” says Dr. Doga. “Our goal is to design quantum algorithms that can find how to predict protein structures as realistically as possible.”